rJava, rCharts and R code to display GC data
July 5, 2014 Leave a comment
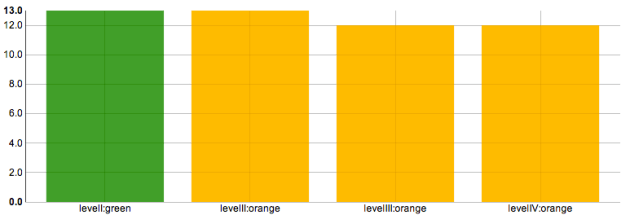
These are the steps I follow to display GC activity data using a nvd3 discrete bar chart.
Call Java class using rJava
gctypes <- .jcall(realtimegcdataobserver ,"Ljava/util/List;","getGCTypes")
Create an empty data frame to hold the data
I get the type of the GC algorithm, GC count and time from JMX. I have yet to explore the last two values.
gcdata <- function(){
df <- data.frame(
GCType=character(),
Count=character(),
Time=character(),
stringsAsFactors=FALSE)
print(df)
return(df)
}
Iterate over the list of beans
Call appropriate methods and fill up the empty data frame.
I massage the data using the last two lines but don’t know any elegant way to accomplish this.
emptygcdata <- gcdata() gctypedetails <- sapply( gctypes, function(item) rbind(emptygcdata, as.data.frame(c(GCType=item$getName(),Count=item$getL(),Time=item$getM())))) gctypedetails <- data.frame(gctypedetails) gctypedetails <- data.frame(matrix(unlist(gctypedetails)))
matrix.unlist.gctypedetails..
1 PS Scavenge
2 16
3 22
4 PS MarkSweep
5 0
6 0
emptygcdata <- gcdata()
before <- 0
after <- 2
repeat
{
if (after >= nrow(gctypedetails))
break;
emptygcdata <- rbind(emptygcdata, data.frame(GCType =gctypedetails[before + 1,1], Count =gctypedetails[before + 2,1], Time=gctypedetails[before + 3,1]))
before <- after + 1;
after <- after + 2;
}
GCType Count Time
1 PS Scavenge 16 22
2 PS MarkSweep 0 0
nvd3 using rCharts
p2 = nPlot(x = "Time", y = "Count", data = emptygcdata, type = "discreteBarChart")
p2$chart(
color = "#! function(d){
var ourColorScale = d3.scale.ordinal().domain(['PS MarkSweep','PS Scavenge']).range(['green','purple']);
return ourColorScale(d.GCType);
}!#")