R Reference classes
June 9, 2017 Leave a comment
A pure OO approach and a functional representation of it are at loggerheads. That is evident when one tries to adopt an OO approach using a powerful functional language. That is my personal opinion.
R has many Object-oriented features built into it.
R has three object oriented (OO) systems: [[S3]], [[S4]] and [[R5]].
Reference classes are one such feature.
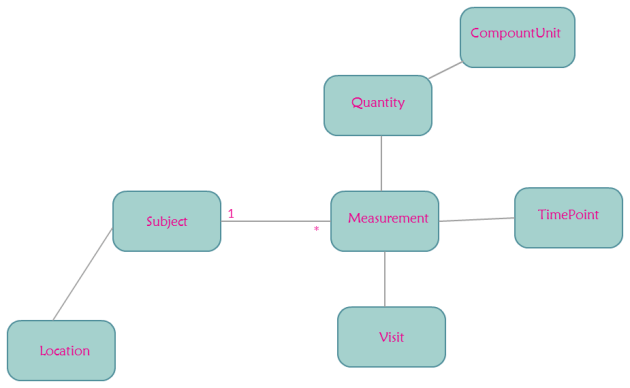
Let us consider this data. The id is that of a Subject who is in
a room where monitoring equipment gathers some data. There are several visits to gather this data.
id visit room value timepoint 14 0 bedroom 6 53 14 0 bedroom 6 54 15 0 bedroom 2.75 56
The idea that this code is based on is from Martin Fowler’s book Analysis Patterns Reusable Object Models. The chapter on Observations and Measurements has a diagram roughly equivalent to the
one shown at the top.
The code is lightly tested several times but without unit tests.
library(plyr) library(dplyr) library(purrr)
CompoundUnit <- setRefClass("CompoundUnit",
fields = list(micrograms = 'numeric',
cubicmeter = 'numeric'))
Location <- setRefClass("Location",
fields = list( room = 'character'),
methods=list(getlocation = function(){
room
},
summary = function(){
paste('Room [' , room , ']')
}))
library(objectProperties)
# An Enum which could have behaviour associated with it.
# This is convoluted but the only way I know to represent constants and validate them.
#
###############################################################################
MeasurementVisitEnum.gen <- setSingleEnum("MeasurementVisit",levels = c('0', '1', '2'))
par.gen <- setRefClass("Visit",
properties(fields = list(visit = "MeasurementVisitSingleEnum"),
prototype = list(visit =
new("MeasurementVisitSingleEnum",
'0'))))
What is the significance of this convoluted code ?
It restricts the values that are set to 0.1 and 2. It is like the Java enum
But this is not strictly a requirement here. It is just that there is a facility to identify erroneous data if we need it.
> MeasurementVisitEnum.gen par.gen visits visits$visit visits$visit visits$visit visits$visit <- as.character(3)
Error in (function (val) :
Attempt to set invalid value on 'visit': value '3' does not belong to level set
( 0, 1, 2 )
TimePoint <- setRefClass("TimePoint",
fields = list(time = 'numeric'))
Quantity <- setRefClass("Quantity",
fields = list(amount = "numeric",
units = CompoundUnit))
Measurement encapsulates the quantity, the time point and the visit number. So, for example, during visit 0, at this time point the quantity was observed. This type of encapsulation in the true spirit of OO has its
disadvantages as we will see later.
Measurement <- setRefClass("Measurement",
fields = list(
quantity = "Quantity",
timepoint = "TimePoint",
visit = "Visit"),
methods=list(getvisit = function(){
visit$visit
},getquantity = function(){
quantity
})
)
Subject <- setRefClass("Subject",
fields = list( id = "numeric",
measurement = "Measurement",
location = "Location"),
methods=list(getmeasurement = function()
{
measurement
},
getid = function()
{
id
},
getlocation = function()
{
location
},
summary = function()#Implement other summary methods in appropriate objects as per their responsibilities
{
paste("Subject summary ID [",id,"] Location [",location$summary(),"]")
},show = function(){
cat("Subject summary ID [",id,"] Location [",location$summary(),"]\n")
})
)
LongitudinalDatum is the class LongitudinalData inherits from. This inheritance is shown as an example. Not all methods that should belong in the super class are properly added. There are many methods in the sub class that can be moved a level up.
subsummary in the super class can be called from the sub class. The line if( subject(x) == id){ in the sub class LongitudinalData calls this super class method.
LongitudinalDatum datum
measurements <<- list()
load(datum)
},load = function( df ){
by(df, 1:nrow(df), function(row) {
visits <- par.gen$new()
visits$visit <- as.character(row$visit)
u <- CompoundUnit$new( micrograms = 1,
cubicmeter = 1 )
q <- Quantity$new(amount = row$value,
units = u )
t <- TimePoint$new(time = row$timepoint)
m <- Measurement$new(
quantity = q,
timepoint = t,
visit = visits)
l <- Location$new( room = as.character(row$room))
s <- Subject$new( id = row$id,
measurement = m,
location = l)
measurements <<- c( measurements, s )
})
},
getmeasurementslength = function(){
length(measurements)
},
findsubject = function( id ){
result % map(., function(x) {
if( subject(x) == id){
result <<- x # Warning message is benign for this example. result
#cannot be a class state. It is really local.
}
}
)
result
},
visit = function( sub,v ){
measurementsvisit % map(., function(x) {
m <- x$getmeasurement()
if (m$getvisit() == v && x$getid() == sub$getid() ){
measurementsvisit <<- c(measurementsvisit,x)
}
}
)
list(visit = measurementsvisit )
}
},
room = function( t, room ){
if( length( t) == 0 ){
c('NA')
}else{
measurementsvisitroom % map(., function(x) {
if( x$getlocation()$getlocation() == room )
measurementsvisitroom <% map(., function(y) {
if (x$getid() == y$getid() ){
m <<- x$getmeasurement()
summaries <% summary
}
},subjectsummary = function( subject ){
filteredmeasurements <-
keep(measurements, function(x){
x$getid() == subject$getid()
})
groupedmeasurements % lapply(function(x){
m <% rbind_all()
dataColumns <- c('amount')
ddply(groupedmeasurements,c('visit','location'),function(x)
colSums(x[dataColumns]))
}
)
)
How does this work ?
The data is loaded into an object hierarchy in the load function. I did observe that it was slow most probably because my Eclipse StatET for R setup needs more memory.
Since the methods are all encapsulated by the class I am using the reference to call methods. The result of findsubject is passed to subjectsummary because I am piping the result of one method to the next.
ld <- LongitudinalData$new() out % ld$subjectsummary() print(out)
So here the result of findsubject(14) is passed as the first parameter when visit(0) is called. 0 becomes the second parameter.
out % ld$visit(0) %>% ld$room("bedroom")
The final result from this pipeline is whatever is returned by the last method room("bedroom").
I would like to reassert that this is just one way of combining multiple methods using Reference classes. There are much more powerful functional approaches that don’t require this many lines of code. This example illustrates a particular Object-oriented approach.
Flattening the Reference classes
The OO hierarchy here does not seem to be malleable when used with some R packagea like dplyr. Try as I may, I cannot coerce the Reference classes into a R data frame and pipe it through stages using dplyr. Remember I want to use functions like map and filter to get the data out of these reference clasees in a shape that I want.
So I abandon my OO approach and flatten the objects and create a data frame. Now I get back the data in the shape I want.
groupedmeasurements %
lapply(
function(x){
m <% rbind_all()
This is how one gets the following output.
out % ld$subjectsummary()
print(out)
visit location amount 0 bedroom 12.00 0 dining room 2.75 0 living room 2.75 0 room 5.50 0 tv room 2.75 1 room 2.75
Conclusion
This exercise has not helped me determine in which context R’s Reference classes are specifically used. The other OO systems like S3 and S4 may be more useful but this article is about RC’s. Why should I flatten my object hierarchy to reshape my data in a convenient way ? There may be specialized R packages that use the OO approach and expose API’s but I am not aware of them. So at this time I understand that there is a dichotomy between RC’s and the powerful functional approach. I personally like to use the functional programming paradigm when dealing with data.