Notes about Machine Learning fundamentals
April 26, 2016 Leave a comment
I have decided to try a different tack in this post. Gradually as I learn some basic ideas about statistics and Machine Learning I will update this post with code, graphs or procedures used to configure tools. So in a few weeks I will have charted a simple course through the basic Machine Learning terrain. I hope. But these are just basic ideas to prepare oneself to read a more advanced math text.
To be updated … I will add more details in subsequent posts.
Tools
Anaconda based on Anaconda
GraphLab based on GraphLab
ipython based on ipython
The installation process was tortuous because I work in a corporate environment.
Install GraphLab Create with Command Line
The installation is based on dato’s instructions.
Step 1: Ensure Python 2.7.x
Anaconda with Python 2.x installation didn’t complete in my Windows 7 machine due to some access restriction. It couldn’t set this version of Python as the default.
So I installed Anaconda with Python 3.x. GraphLab works with only Python 2.x
In order to create a Python 2.7 environment the command used is
conda create -n dato-env python=2.7 anaconda
This was blocked by my Virus Scanner and I had to coax our security team to update my policy settings to allow this.
Traceback (most recent call last):
File “D:\Continuum\Anaconda3.4\Scripts\conda-script.py”, line 4, in <module>
sys.exit(main())
File “D:\Continuum\Anaconda3.4\lib\site-packages\conda\cli\main.py”, line 202,
in main
args_func(args, p)
File “D:\Continuum\Anaconda3.4\lib\site-packages\conda\cli\main.py”, line 207,
in args_func
args.func(args, p)
File “D:\Continuum\Anaconda3.4\lib\site-packages\conda\cli\main_create.py”, li
ne 50, in execute
install.install(args, parser, ‘create’)
File “D:\Continuum\Anaconda3.4\lib\site-packages\conda\cli\install.py”, line 4
20, in install
plan.execute_actions(actions, index, verbose=not args.quiet)
File “D:\Continuum\Anaconda3.4\lib\site-packages\conda\plan.py”, line 502, in
execute_actions
inst.execute_instructions(plan, index, verbose)
File “D:\Continuum\Anaconda3.4\lib\site-packages\conda\instructions.py”, line
140, in execute_instructions
cmd(state, arg)
File “D:\Continuum\Anaconda3.4\lib\site-packages\conda\instructions.py”, line
55, in EXTRACT_CMD
install.extract(config.pkgs_dirs[0], arg)
File “D:\Continuum\Anaconda3.4\lib\site-packages\conda\install.py”, line 448,
in extract
t.extractall(path=path)
File “D:\Continuum\Anaconda3.4\lib\tarfile.py”, line 1980, in extractall
self.extract(tarinfo, path, set_attrs=not tarinfo.isdir())
File “D:\Continuum\Anaconda3.4\lib\tarfile.py”, line 2019, in extract
set_attrs=set_attrs)
File “D:\Continuum\Anaconda3.4\lib\tarfile.py”, line 2088, in _extract_member
self.makefile(tarinfo, targetpath)
File “D:\Continuum\Anaconda3.4\lib\tarfile.py”, line 2128, in makefile
with bltn_open(targetpath, “wb”) as target:
PermissionError: [Errno 13] Permission denied: ‘D:\\Continuum\\Anaconda3.4\\pkgs
\\python-2.7.11-4\\Lib\\pdb.doc’
The last line shown above is what I presume was blocked by the Virus scanner. When the logs were shown to the security team who updated the scanner rules.
Learning some of these topics may be difficult if we don’t read a more advanced book. So I am constrained by the lack of deep knowledge of a related math subject.
But a question like this one must be simple. Right ?
Identify which model performs better when you have the intercept, slope and Residual Sum of squares.
No data point is given.One can plot the lines when their intercepts and slopes are known but I don’t know how that helps.
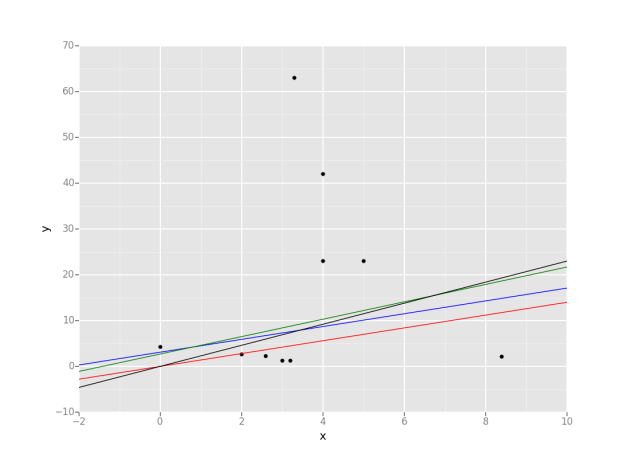
Plot some lines when we know their intercepts and slopes. Data points are random though and are irrevelant at this time.
from ggplot import *
import pandas as pd
data = {'x': [0, 2, 3, 4, 5, 4, 3.2, 3.3, 2.6, 8.4],
'y': [4.2, 2.6, 1.2, 23, 23, 42, 1.2, 63, 2.3, 2.1],
}
df = pd.DataFrame(data)
g = ggplot(df, aes(x='x', y='y')) + \
geom_point() + \
geom_abline(intercept=0, slope=1.4, colour=&quot;red&quot;) \
+ geom_abline(intercept=3.1, slope=1.4, colour=&quot;blue&quot;) \
+ geom_abline(intercept=2.7, slope=1.9, colour=&quot;green&quot;) \
+ geom_abline(intercept=0, slope=2.3, colour=&quot;black&quot;)
print(g)